-
长期载人飞行和载人登月已成为21世纪航天发展的热点。复杂的空间辐射和微重力环境是影响近地轨道飞行中影响航天员身体健康的主要因素。随着今后深空探索频率和滞留时间的增加,地球磁场、大气层及有限的飞行器载荷对宇宙射线的屏蔽效果将逐渐弱化,因此宇宙射线对航天员的影响将逐渐凸显,亟需关注微重力环境下宇宙射线对航天员带来的不良影响。
已有研究[1]发现,航天员近地轨道飞行10~14天后红细胞总量累计下降10%~15%,血红蛋白含量下降12%~13%,出现血浆容量、网织红细胞和血红蛋白减少,异形红细胞增多等血液学变化,航天医学家称之为“航天贫血症”。但美国国家航空航天局(NASA)的研究数据却表明,国际空间站停留6个月以上的宇航员在已经适应了微重力的情况下不会出现严重贫血的情况[2]。因此,“航天贫血症”的发生机制还有待进一步研究。
红系分化是骨髓造血干细胞分化为多向祖细胞,再不断增殖逐步分化为红系祖细胞、红系前体细胞,最后发育为成熟红细胞并释放到外周血中的过程。红系各阶段细胞增殖与分化、生长与凋亡之间的平衡需要多种细胞因子共同调控完成[3]。促红细胞生成素(EPO)能诱导EPO受体(EPOR)形成二聚体,是刺激骨髓红系祖细胞增生、诱导红系分化和促进红细胞成熟的主要因子。磷酸肌醇3激酶(PI3K)是催化磷脂酰4, 5二磷酸(PIP2)转变为磷脂酰3, 4, 5三磷酸(PIP3)从而激活PI3K-AKT通路,进而调控细胞增殖、凋亡等生命活动的一个激酶复合物。PI3K蛋白与激活的EPOR结合[4],进一步激活下游调节细胞周期、生存、分化和代谢的AKT蛋白,也激活一系列红系相关基因,如内源性GATA-1、红系Krüppel样因子(EKLF)、β-珠蛋白(β-globin)等的表达[5]。因此,造血干细胞凋亡与增殖抑制可能造成红系分化障碍。
K562红白血病细胞系[6]能够被羟基脲、阿糖胞苷、hemin等化合物诱导红系分化[7],呈现和正常红细胞相似的表面抗体和血红蛋白合成等表型特征,近年来已被广泛应用于红系分化相关的机制研究[8-9]。我们以前期实验确定的40 μM氯化高铁血红素诱导K562细胞作为红系分化模型,并对该模型进行地面微重力效应模拟及X射线辐照处理,研究二者对红系分化的影响及相关机理。
-
人髓性白血病细胞系K562购自上海富衡生物科技有限公司,RPMI1640培养基购自美国Gibco公司、胎牛血清购自民海公司,双抗(青、链霉素)、CCK-8购自北京Solarbio公司,CD235a抗体、Trizol裂解液购自美国Invitrogen公司,细胞凋亡检测试剂盒购自美国BD公司,cDNA反转录试剂盒购自美国Thermo Scientific公司,2×SYRB Green PCR Master Mix购自美国ABI公司,qPCR引物由武汉天一辉远生物科技公司合成,联苯胺购自天津光复,30%过氧化氢购自上海沪试,亚铁氰化钾购自上海中秦,冰乙酸购自天津富宇,氯化高铁血红素购自北京Solarbio公司,3-MA购自大连美仑公司。
-
X射线生物辐照仪(X-RAD225,PXI公司)、流式细胞仪(Aminis,德国Merck Millipore公司)、酶标仪(Epoch,美国Biotech公司)、荧光正置显微镜(BX-53,德国Olympus公司)、荧光定量PCR仪(StepOnePlus,美国Applied Biosystems公司)。
-
K562细胞常规培养使用体积分数为10%胎牛血清的RPMI1640培养基培养,于体积分数为5%CO2培养箱中37 ℃饱和湿润环境下培养。处于对数生长期的K562中加入氯化高铁血红素(40 μmol/L)诱导作为红系分化模型以进行后续实验。
-
本实验采用细胞回转器来模拟微重力,其原理是样品在电机的带动下不停地绕水平轴匀速旋转,适当转速下的流体剪切力与自由落体相抵消,形成模拟微重力环境。回转器的半径是3 cm,采用转速30 r/min。实验采用X射线辐照仪进行辐照,剂量率为1.15 Gy/min。实验分为5组:空白模型组(Ctr组);1.0 Gy照射组(R组);微重力处理组(M组);1.0 Gy辐照+微重力联合处理组(R+M组);3-MA(PI3K抑制剂)+1.0 Gy辐照+微重力联合处理组(R+M+3-MA组)。Ctr组不经过任何照射和微重力处理;R组仅接受1.0 Gy照射处理;M组仅进行3 h微重力处理;R+M组先模拟微重力处理3 h后,接受1.0 Gy辐照。PI3K抑制剂3-MA的药物处理浓度为5 mmol/L。
-
收集细胞,离心弃上清,加入新鲜培养基300 μL重悬,铺于96孔板中3孔,每孔100 μL,每孔再加入CCK8溶液30 μL,37 ℃继续孵育4 h,终止培养,用酶标仪检测各孔的490 nm吸光度,以空白孔(不接种细胞,其余操作一致)调零,细胞存活率%=实验组A均值/对照组A均值×100%。
-
收集细胞后,离心后弃上清,重悬于250 μL PBS中,加入联苯胺溶液14 μL,30%过氧化氢1 μL,避光孵育2 min,后加入1 μL亚铁氰化钾,孵育10 min。重悬,弃上清,加入10 μL PBS重悬,于显微镜下拍照计数500个细胞,深色为含有血红蛋白细胞的阳性细胞,不着色的为阴性细胞,阳性细胞率%=阳性细胞数/细胞总数×100%。
-
收集细胞,预冷的PBS洗两次,重悬于90 μL PBS中,加10 μL一定比例稀释好的抗体CD235a(1:500),避光孵育30 min。PBS洗细胞3次,重悬于100 μL PBS,用流式细胞仪收集数据,用IDEAS软件分析流式数据。
-
收集细胞,预冷的PBS洗2次,重悬细胞于100 μL凋亡试剂盒中的1×染料结合缓冲液,再依次加入5 μL AnnexinV-FITC和5 μL碘化丙啶染液,混匀,室温避光孵育15 min,于流式细胞仪上进行检测。用IDEAS软件分析流式数据。
-
各处理组细胞在24 h收集,离心后弃上清,将Trizol裂解液加入细胞。静置10 min。加入氯仿,震荡15 s,静置5 min。4 ℃,12 000 g离心10 min,弃去上清液,得到RNA沉淀。用无RNASe的水制备的75%乙醇溶液洗涤RNA沉淀,4 ℃,12 000 g离心5 min,弃上清,加入30~50 μl的无RNASe的水。Illumina HiSeQ-4000第二代测序平台用于测序。实验按照标准Protocol执行,总RNA提取后先进行质量评估,然后进行文库构建、文库质量检测和文库测序。
各样品Clean Data均在5.78 G以上,Q30碱基百分比能达到92.34%。HISAT2用于分别将各样品的Clean Reads与参考基因组进行序列对比。从比对结果统计来看,样品的Reads与参考基因组的比对效率在94.98%~96.05%之间,说明测序结果可信。使用FPKM(Fragments Per Kilobase of transcript per Million fragments mapped)作为基因表达水平的指标。使用DESeq2进行两种情况/组的差异表达分析,将Fold Change
$\geqslant 1.5$ 且FDR(False Discovery Rate)<0.05作为筛选差异表达基因的标准。 -
对GATA-1、EPOR、PIK3R2基因表达进行实时荧光定量PCR检测,引物列表见表1。取2 μg的总RNA用于每份cDNA反转录,cDNA合成按照逆转录试剂盒说明书进行。以获得的cDNA为模板,按照Qiagen SYRB Green荧光定量PCR试剂盒说明书进行实时荧光定量PCR。所有结果经Actin内参校正,用△△CT法计算表达量。
表 1 qRT-PCR引物列表
基因名称 引物序列(5'-3') Actin F:GGACTTCGAGCAAGAGATGG R:CTGTACGCCAACACAGTGCT GATA-1 F:CCAAGCTTCGTGGAACTCTC R:ATTGTCAGTAAACGGGCAGG EPOR F:GAGCATGCCCA GGATACCTA R:TCTGCTGCCAGCTTTGAGTA PIK3R2 F:CCTGGCACCTATGTGGAGTT R:CAGTTCTCCCCACCTGATGT -
数值表示为均数±标准差,采用成组设计资料的t检验,SPSS统计软件和GraphPad Prism8绘图软件。
-
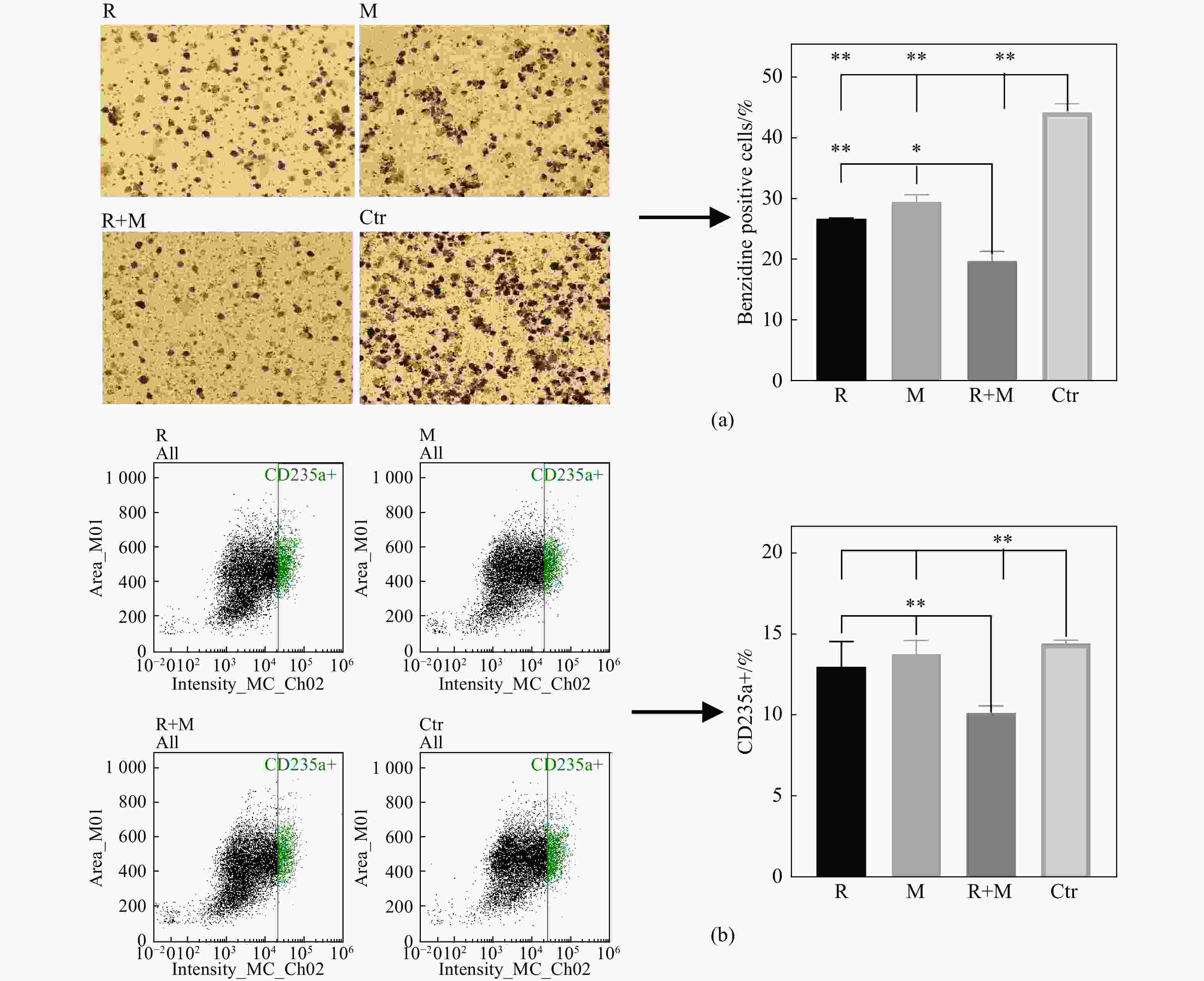
联苯胺染色检测K562细胞中血红蛋白的表达情况,不同处理组的24 h的K562细胞联苯胺染色结果如图1(a)所示,R组、M组、R+M组、Ctr组的联苯胺染色阳性染色率依次为:(26.84±0.11)%, (29.62±1.15)%, (19.94±1.52)%, (44.27±1.38)%。各处理组的联苯胺染色阳性率均与Ctr组有显著性差异(P<0.01),R+M组的阳性率显著低于R组和M组。这些结果说明,辐照及微重力处理后血红蛋白表达显著降低,且二者具有协同作用。
为了验证联苯胺染色结果,同时检测了红系分化过程表面标志蛋白CD235a的表达情况[10][图1(b)]。结果显示,R组、M组、R+M组、Ctr组的CD235a阳性细胞比例依次为:(13.00±1.57)%, (13.77±0.87)%, (10.20±0.43)%, (14.45±0.21)%。各处理组的CD235a蛋白表达下调;R+M组相较于R组及M组,CD235a蛋白表达下调。以上结果说明,辐照及模拟微重力能够抑制K562细胞红系分化,且两者共同的抑制作用大于单独作用。
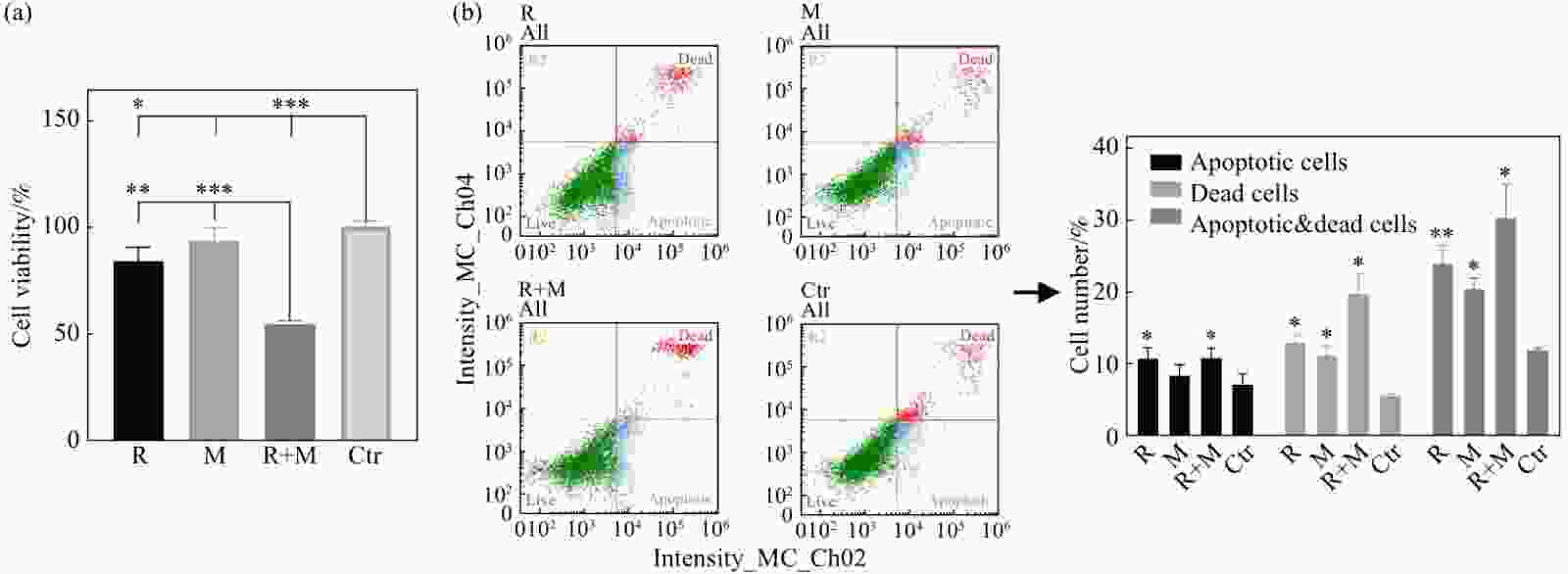
K562细胞作为模型中的红系分化前体细胞,其增殖、凋亡与坏死都可能影响红系分化的结果。各处理组24 h的CCK8结果如图2(a)所示:与Ctr组相比,R+M组的细胞增殖抑制最为显著(P<0.001),R组的增殖抑制也较显著(P<0.05),M组有一定的增殖抑制作用,但无显著性差异(P>0.05)。如图2(b)所示,R组、M组、R+M组、Ctr组K562细胞的凋亡率依次为(10.74±1.55)%, (8.37±1.55)%, (10.81±1.19)%, (7.19±1.45)%;坏死率依次为(12.97±1.08)%, (9.00±1.34)%, (15.40±2.85)%, (8.41±0.31)%;凋亡+坏死细胞比例依次为 (23.93±2.57)%, (17.37±5.39)%, (26.21±7.76)%, (15.60±6.29)%。与Ctr组相比,R组、M组及R+M组的凋亡率及死亡细胞比例均不同程度增加。这些结果说明,X射线辐照及模拟微重力效应能够诱导K562细胞增殖抑制、凋亡及坏死,且二者具有协同效应。
-
通过转录组测定分析得到了各组间差异基因的火山图(图3)。与Ctr组相比,R+M组的差异表达基因数为3514(上调1957,下调1557)。R组与R+M组的差异基因数为15(上调7,下调8),远小于M组与R+M组的差异基因数量236(上调46,下调190)。结果说明,模拟微重力效应及X射线辐照联合作用对K562红系分化模型细胞的基因表达产生较大影响,且主要受X射线辐照处理影响。
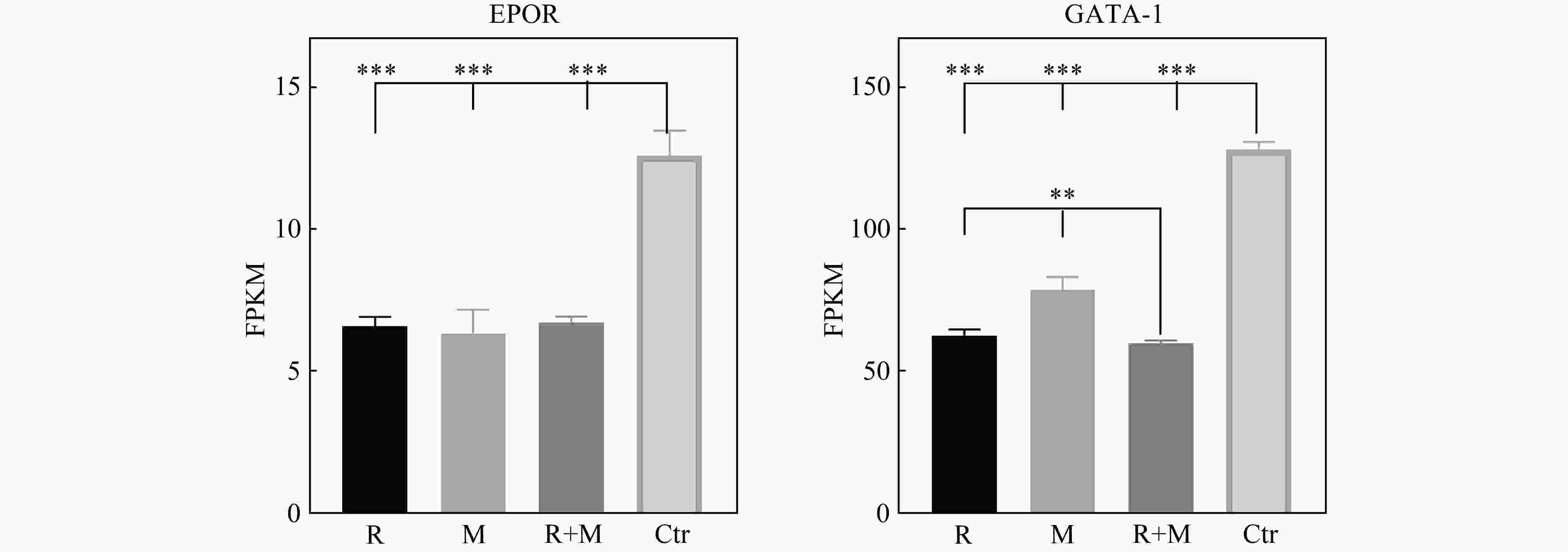
为了确定模拟微重力效应与X射线辐照对红系分化相关因子表达的影响,进一步对比了各处理组24 h红细胞生成素受体EPOR及红系分化特异转录因子GATA-1的基因表达水平,结果如图4所示:R组、M组、R+M组、Ctr组的EPOR的FPKM值依次为6.6089±0.3273, 6.3477±0.8320, 6.744 1±0.199 3, 12.574 5±0.876 3,R组、M组及R+M组相较于Ctr组EPOR的基因表达量均显著下调(P<0.001),R组、M组及R+M组之间无显著性差异(P>0.05)。GATA-1的FPKM值依次为62.957 3±1.962 0, 78.951 5±4.526 4, 60.071 3±1.046 8, 128.018 2±0.681 9。R组、M组及R+M组相较于Ctr组GATA-1的基因表达量均显著下调(P<0.001),R+M组的下调程度大于R组、M组;R+M组与R组无显著性差异,与M组有显著性差异(0.001<P<0.01)。
以上结果说明,模拟微重力效应及X射线辐照可通过下调EPOR及GATA-1的基因表达抑制红系分化,其中X射线辐照对下调GATA-1起主要作用。
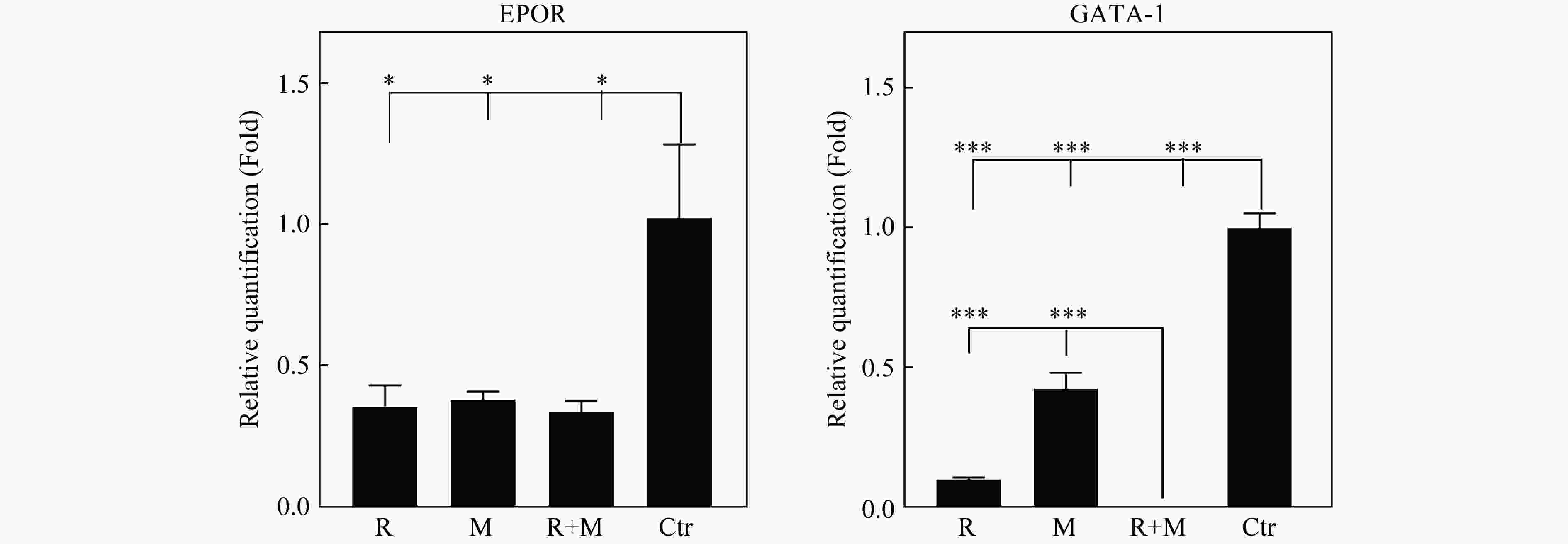
为对转录组结果进行验证,使用qRT-PCR比较了各处理组相对于Ctr组 GATA-1、EPOR基因表达量的差异,结果如图5所示,qRT-PCR的结果与转录组测序各组表达变化一致,说明了转录组测序结果和分析结果的可靠性。
-
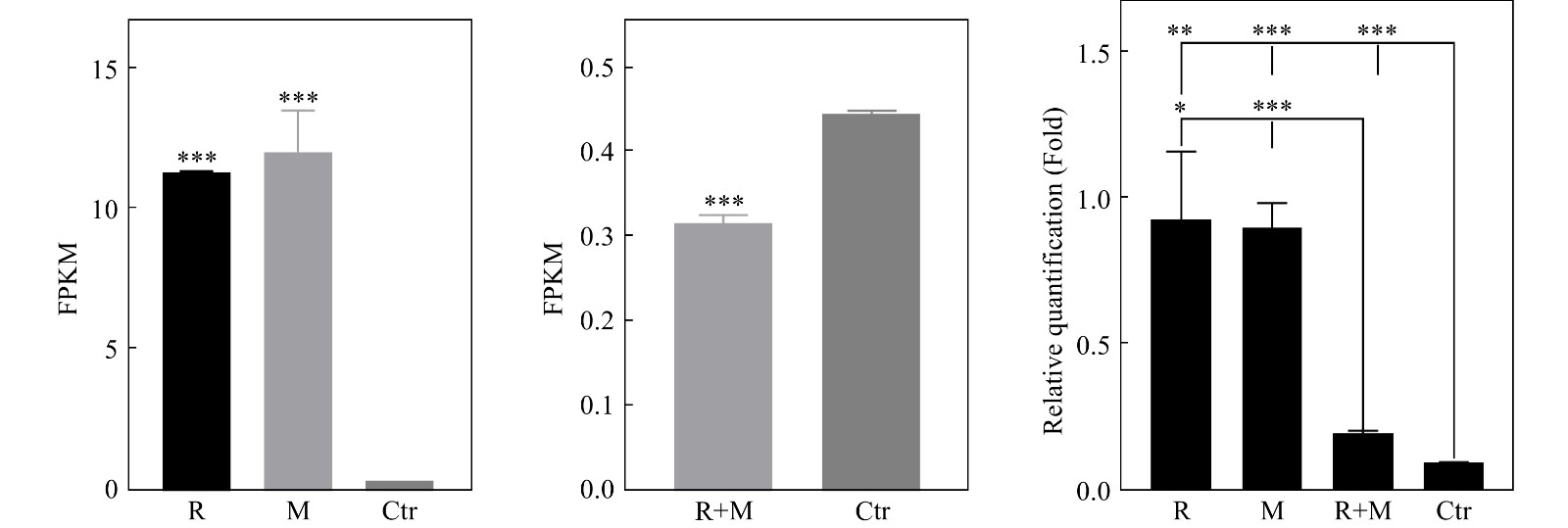
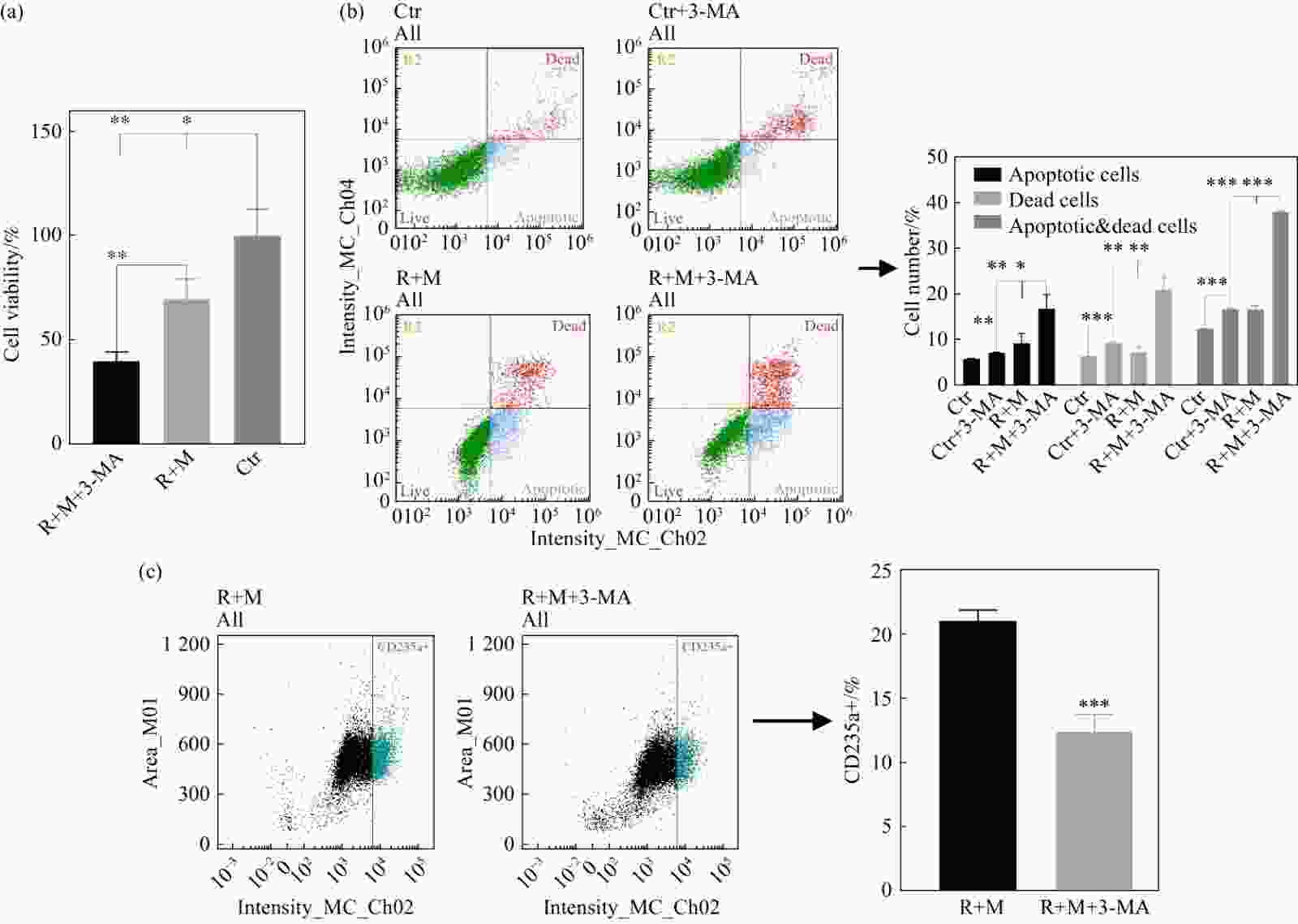
转录组结果显示:R组、M组、R+M组、Ctr组PI3K激酶调控亚基PIK3R2的FPKM值依次为11.254 3±0.044 1, 11.968 0±1.444 8, 0.272 6±0.082 9, 0.443 6±0.003 8。与Ctr组相比,R组及M组的PIK3R2基因表达显著上升(P<0.001),而R+M组的PIK3R2基因表达显著下调(P<0.001)(图6)。PI3K在辐照及微重力处理后表达发生变化,为了验证PI3K在辐照及微重力对K562细胞红系分化模型的影响中发挥的作用,在R+M组中加入PI3K抑制剂3-MA抑制PI3K活性,并检测诱导分化后的K562细胞增殖抑制、凋亡、坏死及红系分化。48 h时,R+M+3-MA组的细胞增殖是R+M组的40.13%,显著受到抑制(P<0.01)。Ctr组、Ctr+3-MA组、R+M组、R+M+3-MA组的凋亡细胞比例依次为(5.97±0.14)%, (7.32±0.11)%, (9.37±2.14)%, (16.87±3.09)%;坏死细胞比例依次为(6.52±0.02)%, (9.49±0.22)%, (7.43±1.19)%, (21.08±2.65)%;凋亡+坏死细胞比例依次为(12.50±0.12)%、(16.81±0.16)%, (16.80±0.70)%, (37.95±0.21)%。R+M组的CD235a阳性细胞比例为(21.17±1.00)%,R+M+3-MA组为(12.80±1.47)%。R+M+3-MA相较于R+M组及Ctr+3-MA组凋亡及坏死细胞比例显著增加,其相较于R+M组凋亡及坏死的增加大于Ctr+3-MA组相较Ctr组凋亡及坏死细胞比例增加。3-MA作用于R+M组细胞后红系分化率降低。以上结果说明,通过抑制PI3K会抑制K562细胞红系分化(图7)。
-
“航天贫血症”能引起人体各系统的生理病理变化,直接影响航天员健康和航天任务的完成,红细胞数量下降是该领域研究的核心问题。红系各阶段细胞增殖与分化的平衡需要多种转录因子共同调控完成[11]。有研究表明,航天员飞行过程中EPO分泌减少会降低外周血红细胞含量[12]。模拟微重力作用于定向祖细胞能明显抑制红系分化和增殖[13]。电离辐射也通过G-CSF/Stat3/BATF通路影响红系分化[14],能造成外周血红细胞减少[15]。但电离辐射与微重力效应的复合作用对红系分化的影响尚不明确,目前尚未见到相关报道。
本实验利用X射线辐照和模拟微重力效应处理K562红系分化细胞模型,红系分化检测结果显示,辐照及微重力效应均能降低红系分化率,且两者能起到协同抑制的作用,其中X射线的效果大于微重力。转录组分析结果显示,微重力与协同处理组的差异基因表达数量远大于辐照与协同处理组的差异基因表达数量,表明辐照在协同处理对基因表达的影响中同样发挥更重要的作用。EPOR及下游因子GATA-1是红系特异性转录因子,能促进红系细胞的分化和成熟。进一步分析转录组数据及qRT-PCR验证发现,辐照及微重力能抑制EPOR及GATA-1的表达,虽然EPOR表达中未显示出协同效应,但其下游基因GATA-1表达显示出协同效应且辐照的影响更大,说明辐照及微重力可以通过抑制EPOR和GATA-1的表达影响K562细胞的红系分化,两者具有协同效应,且辐照的作用大于微重力的作用。
电离辐射和微重力效应均能引起细胞损伤,抑制细胞增殖,诱导细胞凋亡及坏死[15-16]。本研究中各处理组的增殖抑制率、凋亡及坏死率均上升,微重力对细胞损伤的诱导程度低于X射线辐照,两者具有协同效应。X射线辐照及模拟微重力效应对K562细胞的损伤结果[图2(b)]与红系分化结果[图1(a)]负相关,提示细胞生长信号传导途径中的相关因子可能影响红系分化。转录组及qRT-PCR数据显示辐照和微重力处理影响PI3K的调控亚基PIK3R2表达,与对照组相比,辐照、微重力处理后24 h时PIK3R2表达上升,两者联合处理后PIK3R2表达下降,为了进一步研究PI3K与红系分化的相关性,在辐照联合微重力处理前加入PI3K抑制剂3-MA,发现抑制PI3K活性能进一步导致联合处理组细胞增殖活性降低、细胞凋亡与坏死率增高,加重红系分化障碍,说明PI3K在辐照联合微重力造成的红系分化障碍中发挥重要作用。各处理组PI3K不同时间的表达差异及相关机制将在之后的工作中进行探究。
综上所述,X射线辐照及模拟微重力效应对K562细胞红系分化具有协同抑制的作用,不仅可通过特异性分化因子EPOR、GATA-1直接调控红系分化,PI3K调控的生长信号通路相关因子也可能参与其中。转录组分析数据显示各处理组之间还有其它差异表达的基因,其对红系分化的影响机制还有待进一步研究。本实验结果为研究空间血液病的发病机理提供了细胞生物学实验数据,对深空飞行辐射危害的评价及安全措施的制定都具有一定的意义。
Researches on the Synergistic Effect of X-ray Radiation and Simulated Microgravity on Erythroid Differentiation of K562 Cells and Its Mechanism
-
摘要: 使用氯化高铁血红素(hemin)诱导K562细胞分化作为红系分化模型,并对其进行X射线辐照及地面模拟微重力效应处理,研究辐照、微重力及二者的复合效应对红系分化的影响及机制。X射线辐照及模拟微重力效应处理后的联苯胺染色阳性率及CD235a蛋白表达均下调,细胞增殖抑制、细胞凋亡率及坏死率均增高,红系相关转录因子EPOR及GATA-1基因表达下调,且X射线辐照及模拟微重力效应的联合作用效应强于两者单独作用。辐照及微重力联合处理影响细胞中PI3K复合物调控亚基PIK3R2基因表达,加入PI3K抑制剂3-MA后细胞凋亡率及坏死率增高、红系分化率降低。以上结果表明,X射线辐照及模拟微重力效应对红系分化具有协同抑制作用,其机制与红系相关转录因子EPOR、GATA-1及生长信号通路因子PI3K相关。Abstract: In this study, using K562 cells induced by hemin as erythroid differentiation models, the synergistic effects and mechanisms of radiation and simulated microgravity on erythroid differentiation were investigated. Results showed that the positive rates of benzidine staining and the expression of CD235a were down-regulated after treatment with X-ray radiation and simulated microgravity, meanwhile, cell proliferation was inhibited and the rates of apoptosis and necrosis were increased. The expressions of transcription factors related to erythroid differentiation EPOR and GATA-1 were down-regulated. The synergistic effects of X-ray irradiation and simulated microgravity were much stronger than those of the two alone. After irradiation and microgravity combined treatment, PIK3R2 gene expression was down-regulated. After adding PI3K inhibitor 3-MA, the apoptosis and necrosis rate of cells increased, and the erythroid differentiation rates decreased. These results indicate that X-ray irradiation and simulated microgravity treatment have synergistic inhibitory effects on erythroid differentiation, and the mechanism is related to erythroid related transcription factors EPOR, GATA-1 and damage repair pathway factor PI3K.
-
Key words:
- X-ray /
- simulated microgravity effects /
- erythroid differentiation /
- damage repair
-
表 1 qRT-PCR引物列表
基因名称 引物序列(5'-3') Actin F:GGACTTCGAGCAAGAGATGG R:CTGTACGCCAACACAGTGCT GATA-1 F:CCAAGCTTCGTGGAACTCTC R:ATTGTCAGTAAACGGGCAGG EPOR F:GAGCATGCCCA GGATACCTA R:TCTGCTGCCAGCTTTGAGTA PIK3R2 F:CCTGGCACCTATGTGGAGTT R:CAGTTCTCCCCACCTGATGT -
[1] KUNZ H, QUIRIARTE H, SIMPSON R J, et al. BMC Hematology, 2017, 17(1): 12. doi: 10.1186/s12878-017-0083-y [2] AFSHIN B, SHAYONI R, HOMER F, et al. PLOS ONE, 2018, 13(7): e0199621. doi: 10.1371/journal.pone.0199621 [3] STAMATOYANNOPOULOS G. Experimental Hematology, 2005, 33(3): 259. doi: 10.1016/j.exphem.2004.11.007 [4] YU R, HAMILTON G, BAKER D, et al. Journal of Vascular Surgery, 2012, 55(6): 76S. doi: 10.1016/j.jvs.2012.03.192 [5] JOKINEN M, G. GAHMBERG C. Nature, 1979, 278(22): 364. doi: 10.1038/278364a0 [6] MOON A M, LEY T J. Blood, 1991, 77(10): 2272. doi: 10.1182/blood.V77.10.2272.bloodjournal77102272 [7] MAHDI T, ALCALAY D, COGNARD C, et al. Biology of the Cell, 1998, 90(9): 615. doi: 10.1016/S0248-4900(99)80019-7 [8] JIN W P, KANG J Y, HAHM J Y, et al. Communications Biology, 2020, 3(1): 462. doi: 10.1038/s42003-020-01186-8 [9] HU C Y, ZHANG H J, FU C B, et al. Chinese Association of Pathophysiology, 2020, 28(6): 2071. doi: 10.19746/j.cnki.issn.1009-2137.2020.06.045 [10] 马艳妮, 巩蓓, 董贺, 等. 基础医学与临床, 2013, 33(5): 557. MA Yanni, GONG Bei, DONG He, et al. Basic & Clinical Medicine, 2013, 33(5): 557. (in Chinese) [11] KOISTINEN P O, RUSKO H, IRJALA K, et al. Med Sci Sports Exerc, 2000, 32(4): 800. doi: 10.1097/00005768-200004000-00012 [12] GAO Y, PING B H, ZHENG L, et al. Chinese Journal of Microcirculation, 2012, 22(02): 24. [13] ZHENG L, LIU J Z, HU Y W, et al. Aviation, Space, and Environmental Medicine 2011, 82(5):A1. [14] 方连英, 王彦, 徐畅, 等. 基础医学与临床, 2017(2): 256. doi: 10.3969/j.issn.1001-6325.2017.02.036 FANG Lianying, WANG Yan, XU Chang, et al. Basic & Clinical Medicine, 2017(2): 256. (in Chinese) doi: 10.3969/j.issn.1001-6325.2017.02.036 [15] SI J, ZHANG J H, GAN L, et al. Oncology Reports, 2020, 44: 303. doi: 10.3892/or.2020.7581 [16] PRASAD B, D GRIMM, STRAUCH S M, et al. International Journal of Molecular Sciences, 2020, 21(24): 9373. doi: 10.3390/ijms21249373 -




 下载:
下载:




 甘公网安备 62010202000723号
甘公网安备 62010202000723号